Preprocess data with SageMaker Processing, train model with Training and record the parameters and accuracy with Experiments
awsmachinelearningSageMaker Experiments is a feature to record parameters and metrics of Processing and Training jobs etc. In this article, I track from preprocessing to learning as a Run in Experiments and confirm that multiple results can be compared.
The full code is on GitHub.
SageMaker Experiments
experiments.Run() creates an Experiment if it hasn’t existed yet and starts a Run. Previously, it was a separate library called sagemaker-experiments, but now it is integrated into sagemaker.
from sagemaker import experiments
experiment_name = f'test-experiment-{now.strftime("%Y%m%d%H%M%S")}'
with experiments.Run(experiment_name, run_name="run1") as run:
preprocess(run)
train(run, {'aaa': 0.4, 'bbb': True})
Doing load_run() and log_parameter() in each script, parameters are recorded in Run.
from sagemaker.experiments import load_run
with load_run() as run:
run.log_parameter("param1", "value1")
Metrics are also recorded in Experiments, but this is recorded in the Trial Component of the training job instead of Run.
metric_definitions=[{'Name': 'test:accuracy', 'Regex': 'test:accuracy=(\d\.\d+)'}],
enable_sagemaker_metrics=True,
SageMaker Processing
If you call a function like the following from SageMaker Studio or locally, resources will be set up and the script will be executed. In this example, running it on my own image, but you can also use images provided by SageMaker such as PySparkProcessor and SKLearnProcessor.
def preprocess(run: experiments.Run):
processor = Processor(
base_job_name=f'preprocess-{run.run_name}',
image_uri=f'{os.getenv("ECR_REPOSITORY_PREPROCESS")}:latest',
role=os.getenv('ROLE_ARN'),
instance_count=1,
instance_type='ml.m5.xlarge',
env={'AWS_DEFAULT_REGION': os.getenv('AWS_DEFAULT_REGION', 'ap-northeast-1')},
)
processor.run(
inputs=[
ProcessingInput(
source=os.getenv('S3_DATA_PATH'),
destination='/opt/ml/processing/input',
# s3_data_distribution_type='ShardedByS3Key',
),
],
outputs=[
ProcessingOutput(
source='/opt/ml/processing/output',
destination=os.getenv('S3_DATA_PATH'),
# s3_upload_mode='Continuous'
),
],
arguments=[
'--xxx', '12345',
],
wait=True,
)
Source objects are downloaded to the path starting from /opt/ml/processing/ specified as inputs, and once you write files to the path specified as outputs, it will be uploaded to S3. By default, all objects are downloaded to each instance, but s3_data_distribution_type=‘ShardedByS3Key’ makes them sharded. Also, if you set s3_upload_mode=‘Continuous’, files will be uploaded continuously instead of at the end of the job.
import argparse
import pandas as pd
from sagemaker.experiments import load_run
def main():
print("- preprocessing started")
parser = argparse.ArgumentParser()
parser.add_argument('--xxx', type=int, required=True)
args = parser.parse_args()
with load_run() as run:
run.log_parameters({'preprocess:arg_xxx': args.xxx})
df = pd.read_csv("/opt/ml/processing/input/raw.csv")
df.iloc[:5,:].to_csv('/opt/ml/processing/output/train.csv')
df.iloc[5:,:].to_csv('/opt/ml/processing/output/test.csv')
if __name__ == "__main__":
main()
SageMaker Training
Similar to Processing, if you call a function like the following, resources will be set up and training will start. Estimators for each framework are provided by SageMaker, but this time I’m running it on my own image.
SageMakerでPyTorchのモデルを学習させる - sambaiz-net
SageMakerでTensorFlowのモデルを学習させる - sambaiz-net
def train(run: experiments.Run, hyperparameters: object = {}):
estimator = Estimator(
base_job_name=f'train-{run.run_name}',
image_uri=f'{os.getenv("ECR_REPOSITORY_TRAIN")}:latest',
training_repository_access_mode='Platform',
role=os.getenv('ROLE_ARN'),
instance_count=1,
instance_type='ml.m5.xlarge',
hyperparameters=hyperparameters,
environment={'AWS_DEFAULT_REGION': os.getenv('AWS_DEFAULT_REGION', 'ap-northeast-1')},
# output_path=,
)
data_path = os.getenv('S3_DATA_PATH')
estimator.fit(
inputs={'training': data_path, 'testing': data_path},
wait=True
)
inputs are placed in /opt/ml/input/data, and Hyperparameters are written to /opt/ml/input/config/hyperparameters.json. Trained models are uploaded by writing to /opt/ml/model. You can refer to these paths with sagemaker_training library. If no output_path is specified, it will be uploaded to SageMaker’s default Bucket, sagemaker-region-account-id .
import os
from sagemaker.experiments import load_run
from sagemaker_training import environment
import pandas as pd
def train(data_path: str, hyperparameters: dict) -> str:
print(f"data_path: {data_path}")
print(f"hyperparameters: {hyperparameters}")
print(f"training data:\n{pd.read_csv(data_path)}")
return 'trained model'
def test(data_path: str, hyperparameters: dict) -> float:
with load_run() as run:
print(f"test data:\n{pd.read_csv(data_path)}")
run.log_parameters({'test:hp_bbb': hyperparameters.get('bbb')})
for epoch in range(1, 10):
run.log_metric(name="test:accuracy", value=hyperparameters.get('aaa', 0.0) / epoch, step=epoch)
def main():
print("- training started")
env = environment.Environment()
print(f"master_hostname: {env.master_hostname}, current_host: {env.current_host}")
model = train(os.path.join(env.channel_input_dirs['training'], 'train.csv'), env.hyperparameters) # /opt/ml/input/data/training
with open(os.path.join(env.model_dir, 'some_model.dat'), 'w') as f: # /opt/ml/model
f.write(model)
test(os.path.join(env.channel_input_dirs['testing'], 'test.csv'), env.hyperparameters) # /opt/ml/input/data/testing
if __name__ == '__main__':
main()
Experiment results
Check experiment results with ExperimentAnalytics.
from sagemaker.analytics import ExperimentAnalytics
experiment_analytics = ExperimentAnalytics(experiment_name)df = experiment_analytics.dataframe()
df['Source'] = df['SourceArn'].apply(lambda x: str(x).split(':')[-1])
print(df[[
"TrialComponentName",
"Source",
"preprocess:arg_xxx",
"test:hp_bbb",
"test:accuracy - Last"
]])
In addition to TrialComponents which have log_parameter() values, there are ones which have processing-job or training-job as Source. By comparing run1 and run2, you can check which one has higher accuracy along with the parameter values.
TrialComponentName Source preprocess:arg_xxx test:hp_bbb test:accuracy - Last
0 train-run2-2023-05-02-16-40-40-901-aws-trainin... training-job/train-run2-2023-05-02-16-40-40-901 NaN NaN NaN
1 test-experiment-20230503012814-run2 nan 12345.0 False 0.8
2 preprocess-run2-2023-05-02-16-36-12-041-aws-pr... processing-job/preprocess-run2-2023-05-02-16-3... NaN NaN NaN
3 test-experiment-20230503012814-run1 nan 12345.0 True 0.4
4 train-run1-2023-05-02-16-33-44-735-aws-trainin... training-job/train-run1-2023-05-02-16-33-44-735 NaN NaN NaN
5 preprocess-run1-2023-05-02-16-28-15-055-aws-pr... processing-job/preprocess-run1-2023-05-02-16-2... NaN NaN NaN
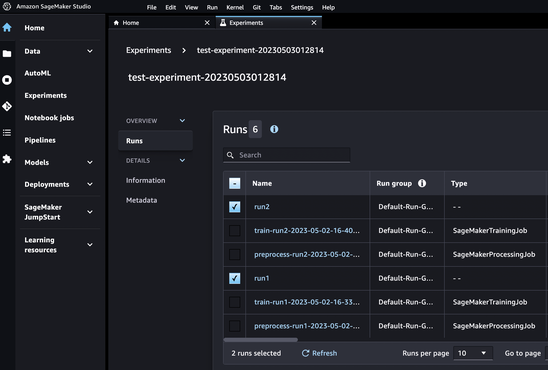
Visualize on SageMaker Studio
Result of experiments can be checked on SageMaker Studio as well.
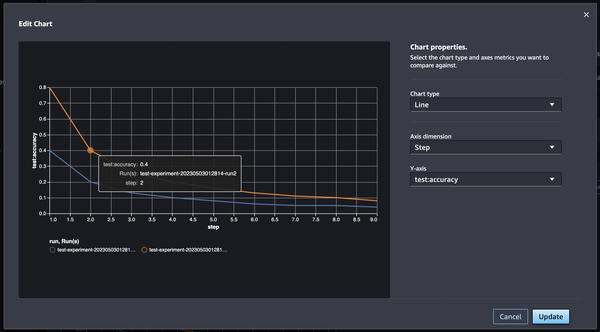
Selecting runs and clicking Analyze, accuracy of each step can be visualized.