EMR Studio is a Jupyter Notebook-based IDE that runs on EMR clusters. While cluster and S3 charges apply, the service itself is free. With EMR Serverless, you can use it easily without having to maintain constantly running clusters.
As a similar service, Athena for Apache Spark is available and while it’s easier to use as an extension of regular Athena, it has some limitations, such as no support for MLlib. Additionally, EMR Studio offers richer features like external IdP authentication and Git integration.
Athena for Apache Spark の Notebook で DataFrame.toPandas().plot() した際の日本語が文字化けしないようにする - sambaiz-net
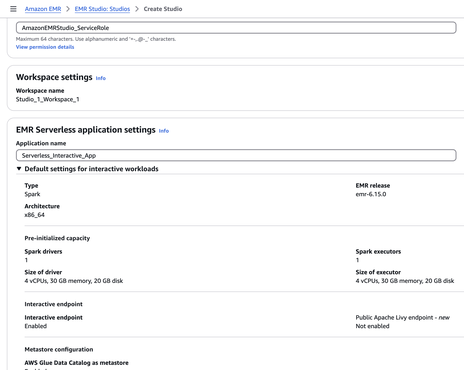
When you create a Studio,
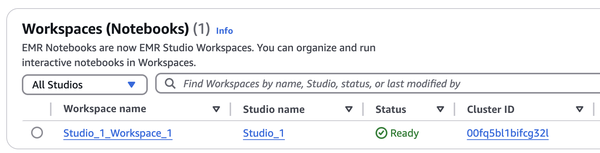
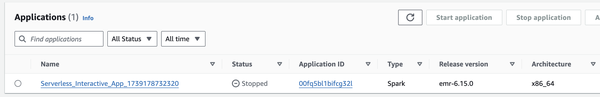
Along with it, EMR Serverless and its associated Workspace (Notebook) are created. The Cluster ID corresponds to the Application ID, which can be confusing since you won’t find it when searching for clusters.
When running interactive workloads on EMR Serverless, kernels that have been idle for more than 60 minutes are automatically terminated from Notebooks. This time limit cannot be changed. Additionally, there is also an auto-stop setting for the Application itself. Even if the Application stops, the Workspace doesn’t disappear.
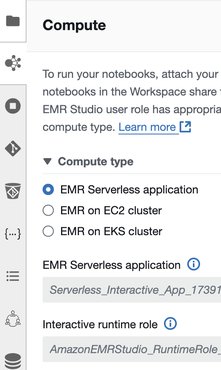
When you configure VPC settings for Studio, you can switch to EMR on EC2 or on EKS. Clusters need to have Spark, Livy, and Jupyter Enterprise Gateway installed. Note that EMR Serverless also has network settings, and changes to Studio settings will not be reflected. While it seems to be able to access S3 even with the default “No network connectivity” setting, it can’t connect to the Internet.
Let’s try running MLlib. By default, the Interactive runtime role only has EMR Studio’s S3 permissions, so you’ll need to grant glue and S3 permissions as needed.
Clustering by k-means method with MLlib of Spark - sambaiz-net
from pyspark.sql import SparkSession
from pyspark.ml.feature import VectorAssembler
from pyspark.ml.clustering import KMeans
spark = SparkSession.builder.appName("test").getOrCreate()
data = [
(0, 1.0, 2.0),
(1, 1.5, 1.8),
(2, 5.0, 8.0),
(3, 6.0, 9.0),
(4, 9.0, 3.0),
(5, 9.5, 2.5)
]
columns = ["id", "x", "y"]
df = spark.createDataFrame(data, columns)
assembler = VectorAssembler(inputCols=["x", "y"], outputCol="features")
vectorized_data = assembler.transform(df)
kmeans = KMeans(k=3, seed=1, featuresCol="features", predictionCol="prediction")
model = kmeans.fit(vectorized_data)
predictions = model.transform(vectorized_data)
predictions.show()
'''
+---+---+---+---------+----------+
| id| x| y| features|prediction|
+---+---+---+---------+----------+
| 0|1.0|2.0|[1.0,2.0]| 1|
| 1|1.5|1.8|[1.5,1.8]| 1|
| 2|5.0|8.0|[5.0,8.0]| 0|
| 3|6.0|9.0|[6.0,9.0]| 0|
| 4|9.0|3.0|[9.0,3.0]| 2|
| 5|9.5|2.5|[9.5,2.5]| 2|
+---+---+---+---------+----------+
'''